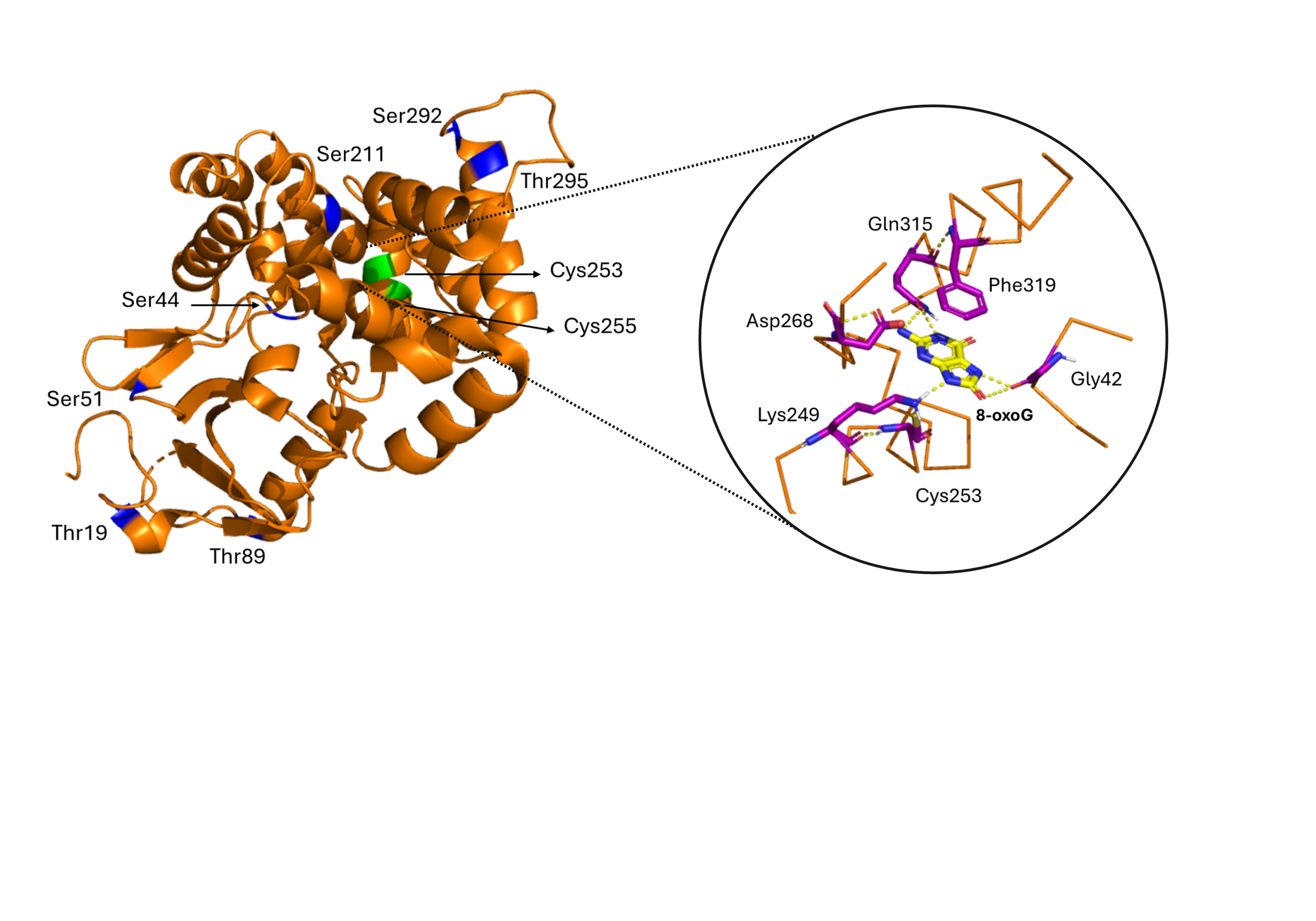DNA Repair Enzymology and Medicinal Chemistry
We study DNA repair enzymes as dynamic regulators of genome stability, transcription and inflammatory signalling. Our work focuses on enzymatic mechanisms that control repair kinetics, pathway choice and product identity, with particular emphasis on Schiff-base chemistry and non-canonical catalytic activities. By dissecting these mechanisms at the biochemical and cellular level, we aim to understand how repair capacity declines with age and disease - and how it can be restored. The tools of choice are small molecules and engineered protein variants.
Medicinal Chemistry
Our Medicinal chemistry efforts for ORCAs encompass the rational design, synthesis and optimization of small molecules. It focuses on engineering the correct acid-base properties (notably pKa tuning) to enable efficient proton abstraction from the Schiff-base intermediate. Structure-based design and SAR studies guide placement of nitrogen functionalities to balance active-site affinity with catalytic competence while avoiding competitive inhibition. Our Medchem efforts also optimize physicochemical and ADME properties to overcome limitations observed with first-generation ORCAs such as TH10785. Special emphasis is placed on organelle targeting, enabling selective delivery to nuclear or mitochondrial OGG1 isoforms. Selectivity profiling and target engagement assays ensure on-target activation without off-target liabilities. Iterative design cycles integrate computational modeling, biochemical kinetics and cellular readouts to refine activity. Scaffold hopping and novel synthetic methods have expanded accessible chemical space beyond nucleobase mimetics. Ultimately, ORCA-focused medicinal chemistry aims to deliver robust chemical probes and drug candidates that enhance and rewire DNA repair in disease-relevant contexts.
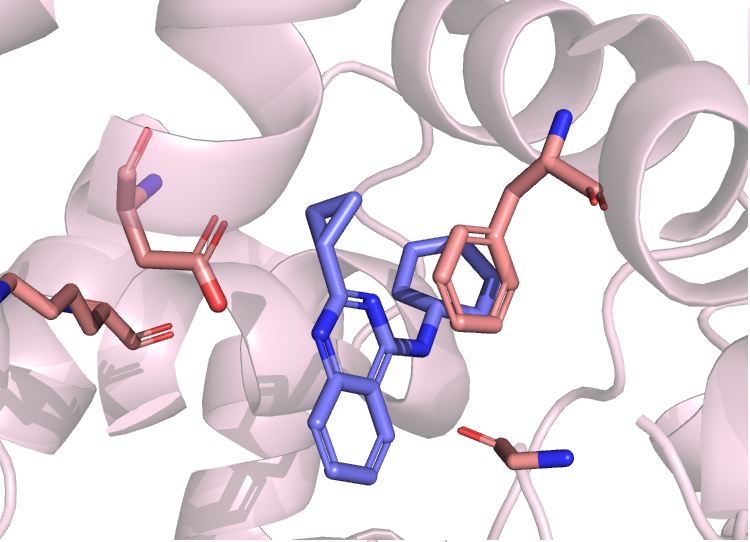
DNA Repair Enzymology
Our protein engineering and enzymology efforts have their origin in OGG1. Now we work on COPs focusing on the rational design and evolution of OGG1 variants with enhanced or rewired catalytic function. It combines structure-guided mutagenesis, QM/MM-informed design, and directed evolution to introduce change-of-function mutations that promote efficient AP-lyase or novel β,δ-elimination activity. Besides next generation protein targets and additional, classical inderstudied, DNA repair enzymes, including the remaining DNA glycosylases, these COPs are profiled through detailed enzymatic kinetics, DNA binding studies and structural biology to map structure-function relationships. Parallel work optimizes expression, folding and resistance to cellular degradation. Delivery strategies are integrated early to ensure intracellular functionality in nuclear or mitochondrial contexts. Selectivity and pathway effects are validated in cellular BER assays and live-cell recruitment studies. Iterative design cycles combine computational modeling with experimental screening to converge on high-performance variants. Ultimately, COPs engineering aims to produce robust, targetable biologics that complement ORCAs by autonomously enhancing and rewiring DNA repair.
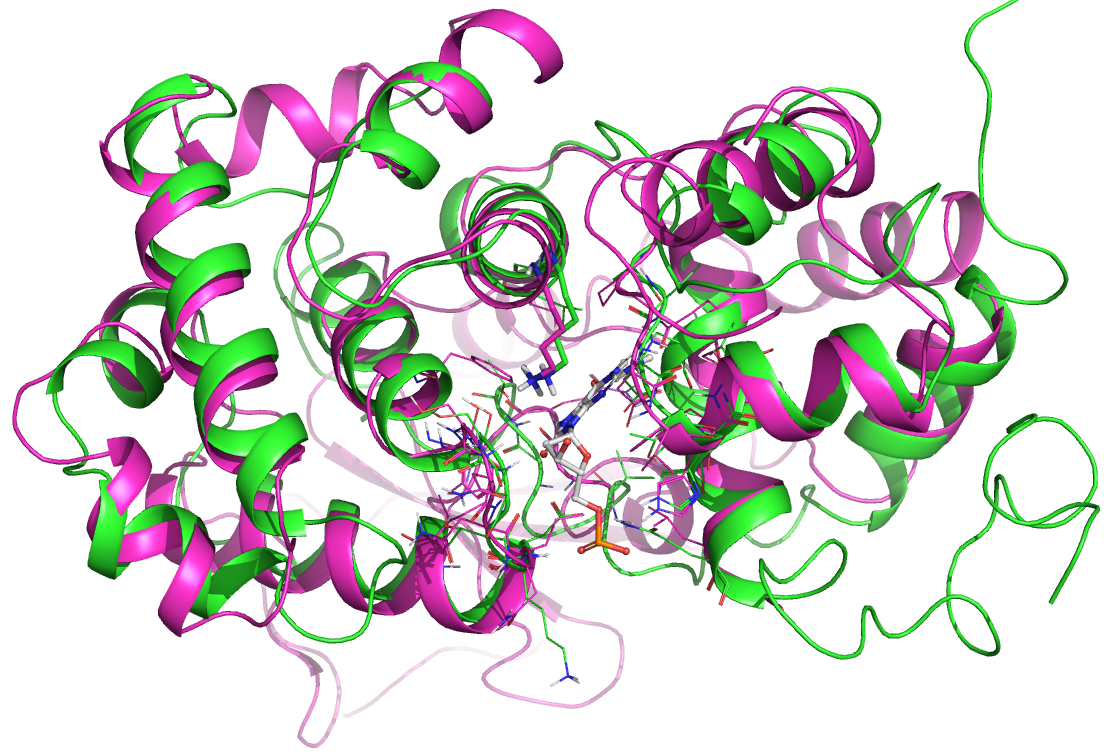
DNA glycosylases
Eleven human DNA glycosylases exist and most of them are considered understudied. In extension of the EUbOPEN consortium and the SGC, we have joined forces with partners in Sweden and Norway to assemble all of them physically. We have developed a platform that includes in vitro and cellular assays for selectivity/activity readout and target engagement. In collaboration with partners at the SGC and Target 2035 we perform large scale screens, solve crystal structures, generate selective antibodies and other reagents and enable investigation of the entire protein family – effectively deorphanizing them for investigation. We donate this setup to the scientific community through protocols within EUbOPEN and Target2035. Ultimately, we thrive to combine our work on ORCAs with these novel targets and enable the studying of DNA repair biology.